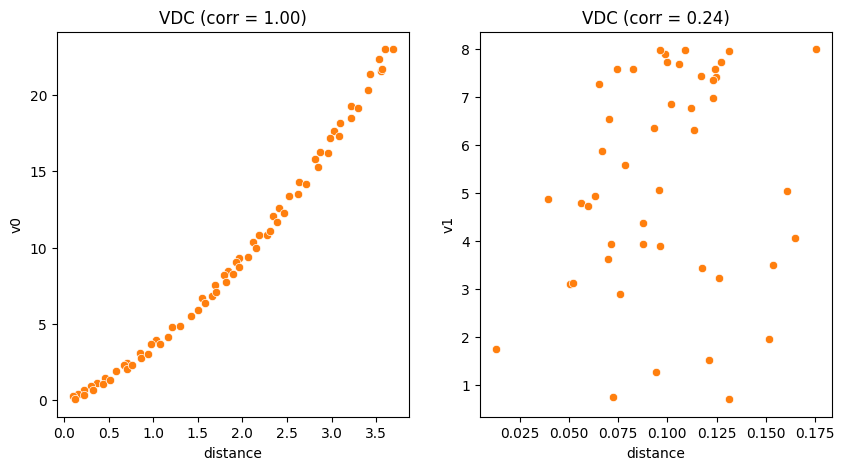
Violation Distance Correlation (VDC)¶
import math
from pyxla import vdc
from pyxla.util import load_data
from pyxla.sampling import HilbertCurveSampler
sphere_sample = {
"name": "Sphere",
"X": HilbertCurveSampler(sample_size=100, dim=1, l_bound=-5, u_bound=5, seed=42),
"F": lambda x: x**2,
"V": [lambda x: x**2 - 2, lambda x: 8 * math.sin(20 * x)]
}
load_data(sphere_sample)
WARNING:root:The Hilbert curve with dimension 1 is just a number line. You are sampling around points on a number line.
vdc(sphere_sample)
WARNING:root:No D file is present, thus, computing the D file... Computing an entire D file can be time consuming. Instead, you can call the function with the keyword argument `compute_D_file` set to `False` to speed up computation, as only the required distances will be calculated.
INFO:root:D file has been loaded to the current sample and is saved to ./Sphere_D.csv
({'vdc_v0': np.float64(0.9988178975694538),
'vdc_v1': np.float64(0.24340888066604996)},
<Figure size 1000x500 with 2 Axes>)